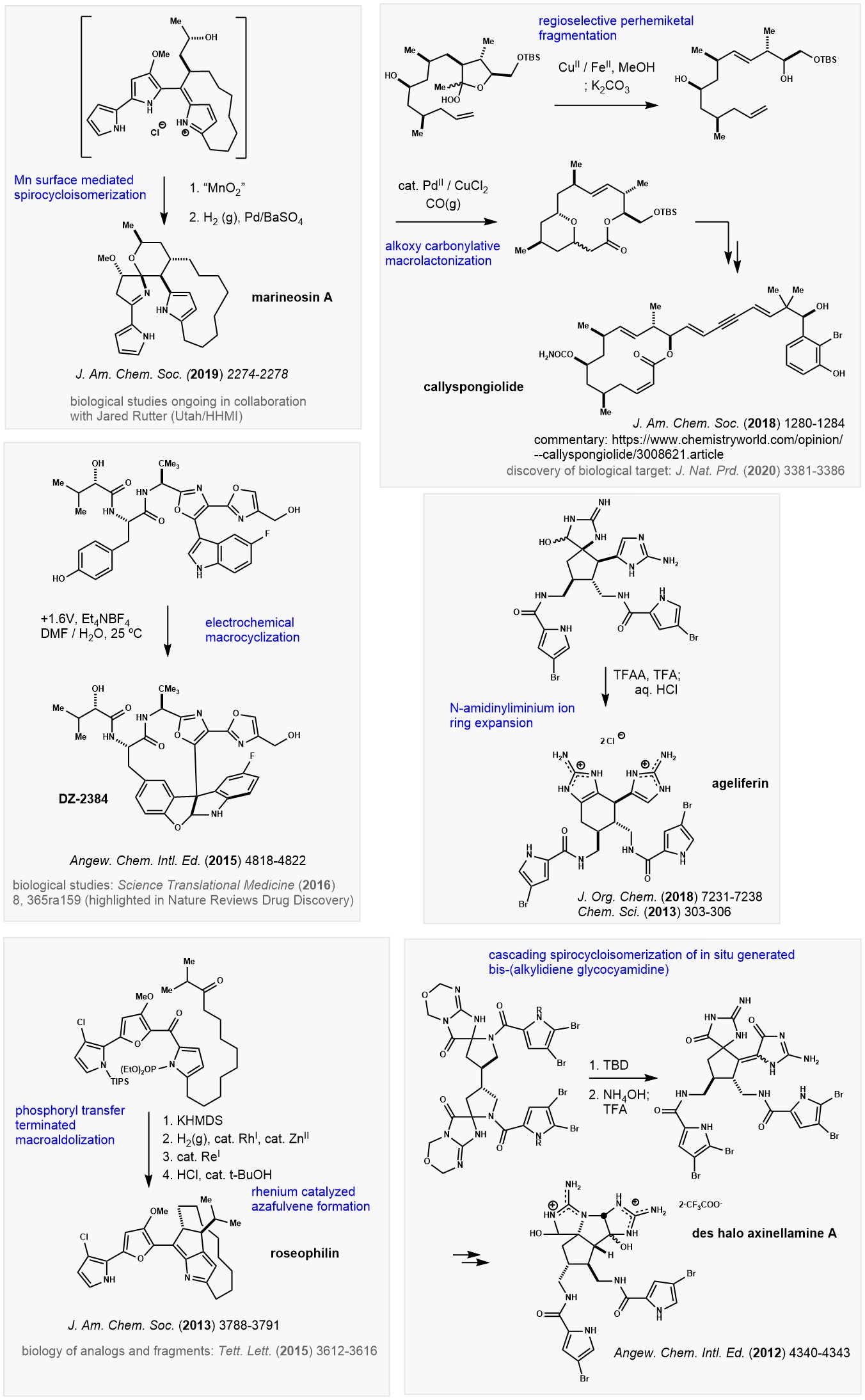
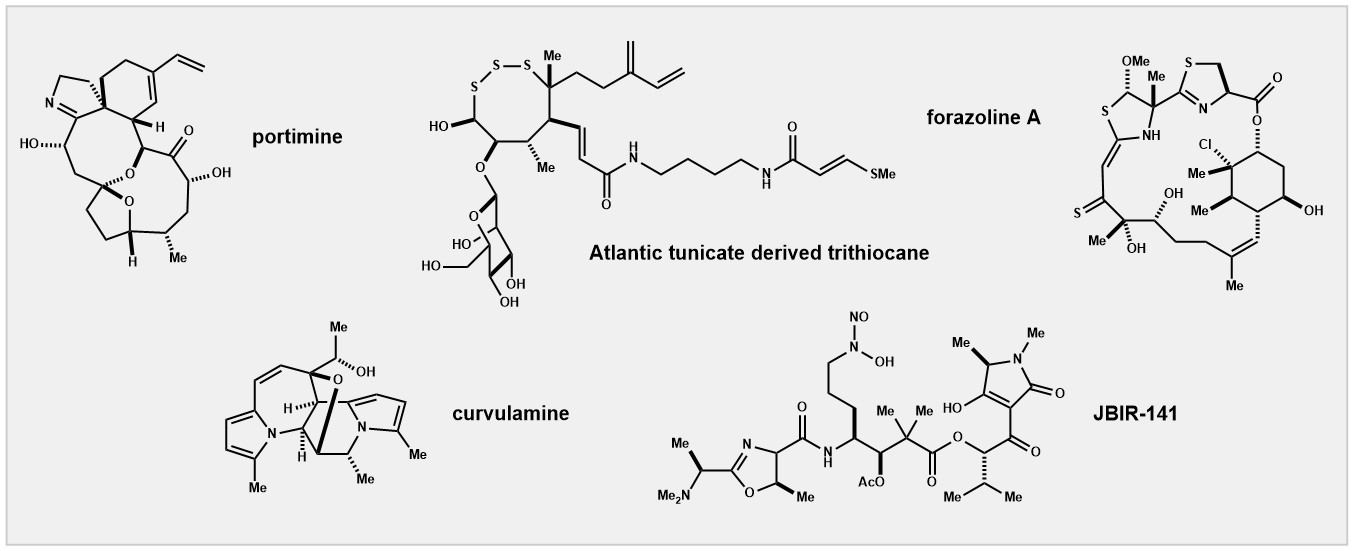
The chemistry and biology of natural products is prominent in the group. We
also study methods to create non-natural molecules – particularly those useful in biological
research. Cellular signaling pathways are a web of protein-protein interactions. Arguably there
is no group of small molecules better suited to probe and perturb those networks than segments
of protein, i.e. peptides. However, in isolation, small peptides generally have poor properties:
they lose structure – they aggregate – they are degraded readily – and they tend not to move
passively through membranes. How can we offset those liabilities while retaining molecular
recognition – and do so systematically?
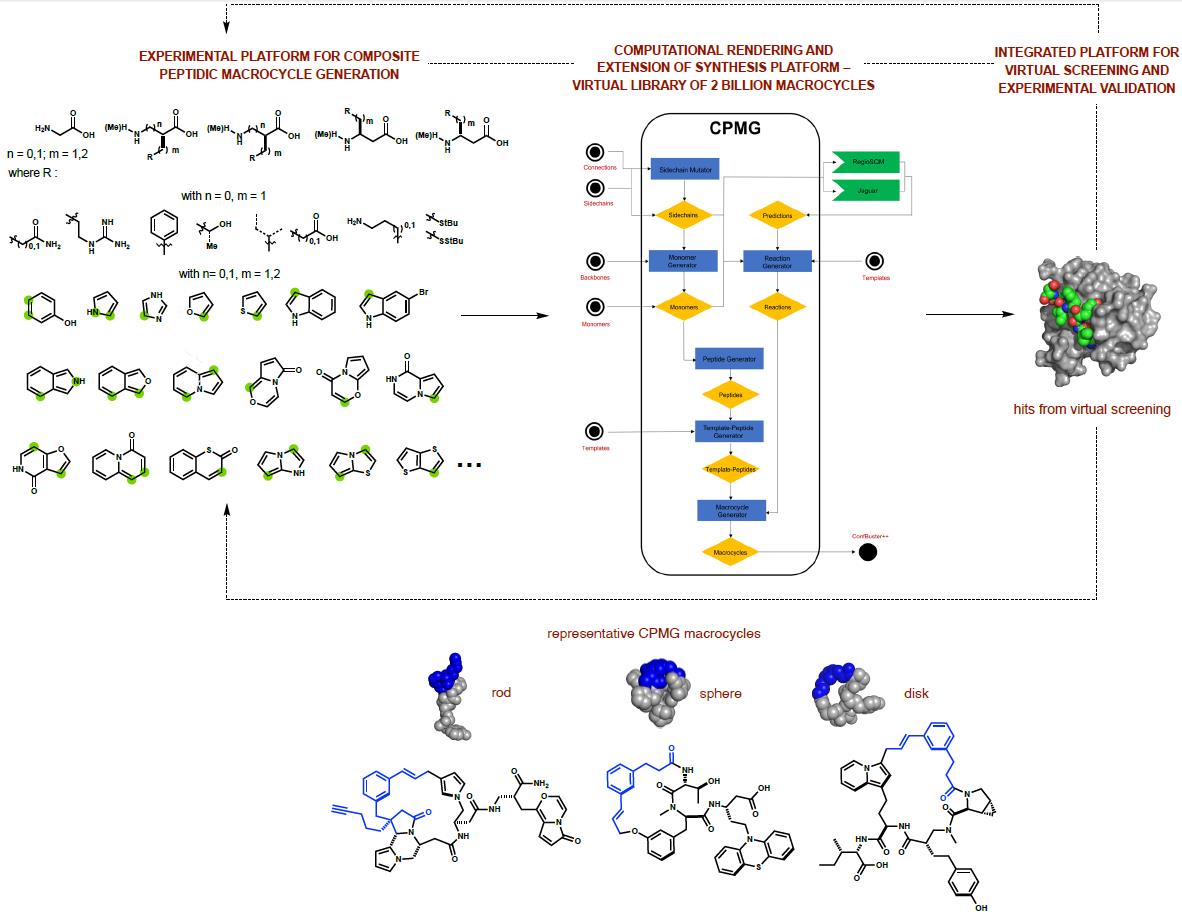
Our approach to this challenge uses constrained, hybrid macrocycles. We have designed
scaffolding reagents that can be integrated into peptide structure to afford diverse ring
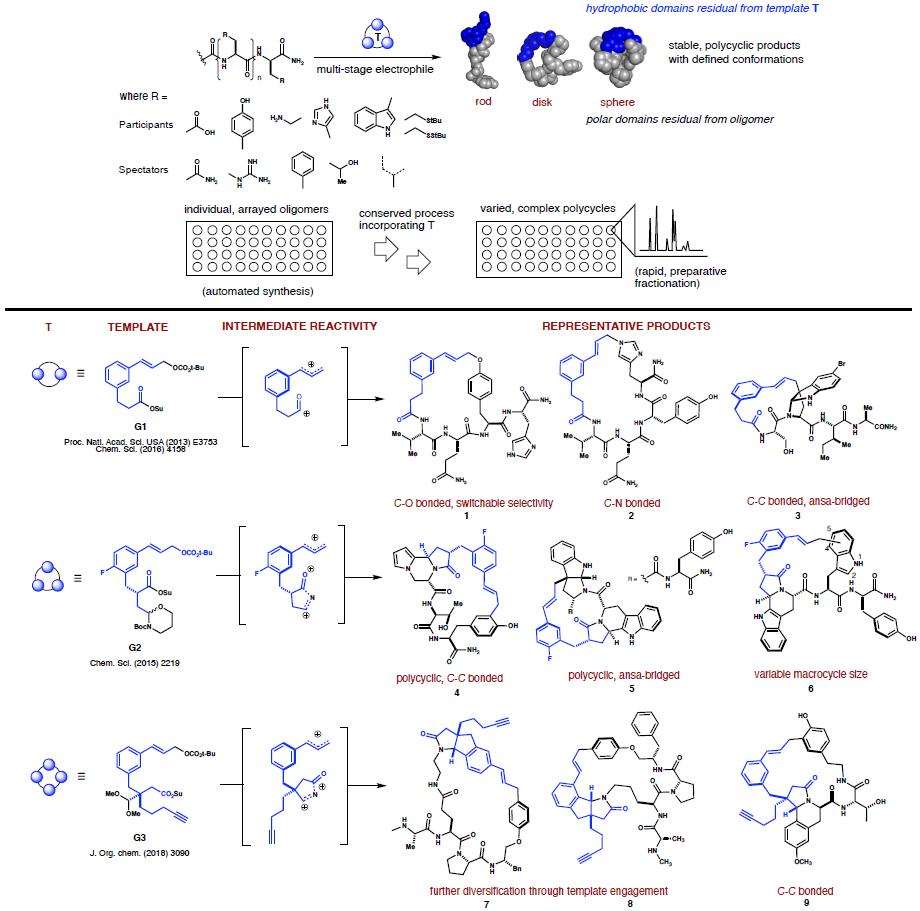
systems having embedded heterocyclic motifs. Our experiments run as processes, wherein templates
G1-G3 are reacted incrementally with unprotected oligomers to form composite products. These
compounds retain molecular recognition elements in the oligomer, yet display that functionality
as part of stable polycyclic structures having improved pharmacological properties. The
methodology allows systematic alteration of product topology by engaging a range of native
peptide functional groups in carbon–heteroatom and carbon–carbon bond-forming reactions.
Templates G1–G3 engage aromatic side chains (including but not limited to phenol, indole, and
imidazole) in Friedel Crafts alkylations, metal-catalyzed allylic substitutions and
N-acyliminium ion-mediated cyclizations. Catalyzed macrocyclization affords C-O or C-N bonded
macrocycles, wherein chemoselectivity is switchable via additives. Macrocyclizations under
acidic conditions generate C-C bonded products via electrophilic aromatic substitutions. These
structures can also incorporate polycyclic motifs via sequential, diastereoselective
N-acyliminium ion cyclizations. G3 is able to itself participate in N-acyliminium ion-promoted
EAS reactions to yield structures such as 7 and 8. Templates
G1 and G2 can generate bridged
endopyrroloindolines such as 3 and 5. New methods and
templates continue to be developed in the
lab.
Based on reactivity patterns observed with G1–G3, a much broader platform
seemed possible, wherein we could process α-amino acid derived oligomers and also those derived
β2- and β3-amino acid monomers, in multiple diastereoisomeric forms. Side chains on those
monomers could harbor diverse, drug-like heterocycles, chosen for their susceptibility to
engagements by G1–G3. End products in that case could have marked property advantages relative
to conventional cyclopeptides. This thought experiment presented a special challenge— possible
outcomes were vast and far outpaced our experimental format. We therefore turned to a
computational rendering of our synthesis platform to systematically assess the scope of reaction
outcomes. We have written a software package called the Composite Peptide Macrocycle Generator
(CPMG). CPMG has generated an in silico library of >2 billion composite macrocycles by
anticipating products from encoded multi-step reaction sequences. We have adapted conformational
search methods to create Confbuster++, which is able to generate three-dimensional conformations
for each macrocycle generated. This positions us to deploy the library in ultra-large scale
virtual screens, with the intent of validating our combined computational/experimental platform
as a tool for discovering unique ligands for all types of structurally characterized proteins.